Table of Contents
Understanding Gaussian Process with Stan in Python
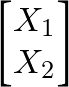
A Gaussian process is a distribution over functions. Instead of assuming a fixed parametric form such as
y = a + bxwe define a prior over all plausible functions and let the observed data update that prior. The key idea is simple: any finite collection of function values has a multivariate normal distribution.
If a latent function is written as
f(x) ~ GP(m(x), k(x, x'))then for observed input points x[1], ..., x[N], the vector
[f(x[1]), ..., f(x[N])]follows a multivariate normal distribution with mean vector determined by m(x) and covariance matrix determined by the kernel k(x, x').
The covariance kernel
The covariance kernel controls what kinds of functions are plausible before seeing the data. A common choice is the exponentiated quadratic kernel, also called the squared exponential or radial basis function kernel:
k(x_i, x_j) = alpha^2 exp(-(x_i - x_j)^2 / (2 rho^2))The parameters have intuitive roles:
alphacontrols the vertical scale of the function.rhocontrols the length scale, or how quickly the function can vary along the input axis.sigmais usually added separately as observation noise.
For noisy observations, the model is
y_i = f(x_i) + epsilon_i
epsilon_i ~ Normal(0, sigma)Equivalently,
y ~ MultiNormal(0, K + sigma^2 I)where K is the covariance matrix built from the kernel.
Why Stan works well for Gaussian processes
Stan is useful here because Gaussian process regression is a Bayesian model with unknown hyperparameters. We usually do not know the correct values of alpha, rho, and sigma, so we place priors on them and infer them from the data.
The tradeoff is computational cost. A full Gaussian process requires Cholesky decomposition of an N x N covariance matrix, which scales roughly as O(N^3). This is fine for small and medium examples, but for large datasets one should consider sparse or approximate Gaussian process methods.
Stan model
The following Stan program uses the exponentiated quadratic covariance function. It adds a small jitter term to the diagonal for numerical stability.
data {
int<lower=1> N;
array[N] real x;
vector[N] y;
}
transformed data {
real delta = 1e-9;
}
parameters {
real<lower=0> alpha;
real<lower=0> rho;
real<lower=0> sigma;
}
model {
matrix[N, N] K;
matrix[N, N] L_K;
alpha ~ normal(0, 1);
rho ~ inv_gamma(5, 5);
sigma ~ normal(0, 0.5);
K = gp_exp_quad_cov(x, alpha, rho);
for (n in 1:N) {
K[n, n] = K[n, n] + square(sigma) + delta;
}
L_K = cholesky_decompose(K);
y ~ multi_normal_cholesky(rep_vector(0, N), L_K);
}Save it as gp_regression.stan.
Python example with CmdStanPy
The Python side only needs to prepare the data, compile the Stan program, and sample from the posterior. Check the current CmdStanPy installation instructions for your environment, because installation details can change. A local sanity check is:
python -c "import cmdstanpy; print(cmdstanpy.__version__)"A minimal reproducible example:
import numpy as np
import matplotlib.pyplot as plt
from cmdstanpy import CmdStanModel
rng = np.random.default_rng(20211202)
N = 40
x = np.linspace(-3, 3, N)
y_true = np.sin(x)
y = y_true + rng.normal(0, 0.15, size=N)
model = CmdStanModel(stan_file="gp_regression.stan")
fit = model.sample(
data={"N": N, "x": x.tolist(), "y": y.tolist()},
chains=4,
iter_warmup=1000,
iter_sampling=1000,
seed=20211202,
)
print(fit.summary().loc[["alpha", "rho", "sigma"]])
posterior = fit.draws_pd(vars=["alpha", "rho", "sigma"])
print(posterior.describe())
plt.scatter(x, y, label="observed", color="black")
plt.plot(x, y_true, label="true function", color="tab:blue")
plt.legend()
plt.show()This model estimates the covariance hyperparameters, but it does not yet generate predictions at new input points. For prediction, add test inputs to the Stan data block and use the conditional multivariate normal formula, or generate posterior predictions in Python using the sampled hyperparameters.
Posterior prediction formula
Let x_train be the observed inputs and x_test be new inputs. Define:
K = k(x_train, x_train) + sigma^2 I
K_star = k(x_train, x_test)
K_test = k(x_test, x_test)The conditional distribution of the latent function at test points is
f_test | y ~ Normal(K_star^T K^{-1} y,
K_test - K_star^T K^{-1} K_star)In implementation, avoid directly computing K^{-1}. Use a Cholesky factor and triangular solves instead. That is both faster and more numerically stable.
Practical checks
For a Gaussian process fitted with Stan, I usually check the following:
Rhatshould be close to1.0foralpha,rho, andsigma.- Effective sample size should not be extremely small.
- Posterior predictions should be smooth at a scale consistent with the learned
rho. sigmashould be plausible relative to the observed noise.- If the sampler reports divergences, inspect priors and rescale the input data.
Input scaling matters. A length scale prior that works for x in [-3, 3] may be inappropriate for x in [0, 100000]. A stable workflow is to standardize or rescale inputs first, fit the model, and then interpret the length scale on that transformed scale.
Summary
A Gaussian process defines a prior over functions through a covariance kernel. Stan lets us infer the kernel hyperparameters instead of fixing them by hand. The basic implementation is compact: build the covariance matrix, add observation noise and jitter to the diagonal, use a Cholesky decomposition, and sample the hyperparameters. For small datasets this gives a clear and flexible Bayesian regression model; for large datasets, the same ideas remain useful, but approximate methods become necessary.
